†equally contributed authors *corresponding author
2026
96. V. H. Truong, D. J. Miller, S. Fatma, Y. Sheng, C. Pindi, M. Ahsan, G. Palermo and E. H. Kellogg. Transposon end recognition and pairing by I-F3 CRISPR-associated transposase. Sci. Adv. 2026, accepted.
95. A. Saha, R. Fregoso Ocampo, J. Wright, D. W. Taylor* and G. Palermo*. Molecular Mechanisms and Biotechnological Applications of CRISPR-Cas12a. Nat. Rev. Mol. Cell. Biol. 2026.
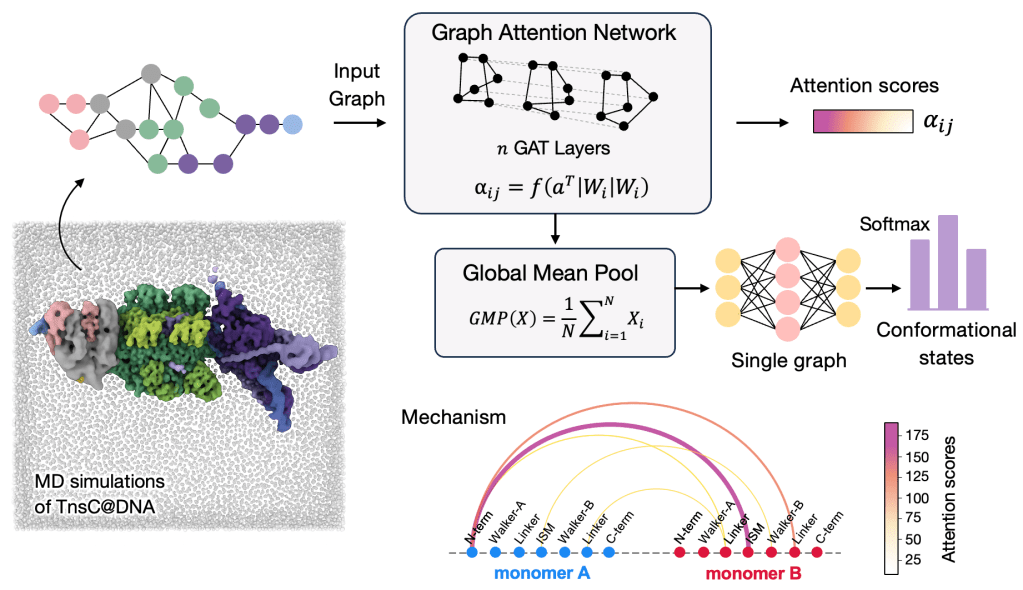
94. M. Ahsan, C. Pindi, S. Sinha, A. C. Patel and G. Palermo*. Graph Neural Networks for Molecular Dynamics Simulations. Curr. Opin. Struct. Biol. 2026, 97, 103238.
93. C. Pindi and G. Palermo*. Computation and deep-learning–driven advances in CRISPR genome editing. Nat. Struct. Mol. Biol. 2026, 33, 203-214 (read PDF)
93. C. Pindi and G. Palermo*. Deep learning and cryogenic electron microscopy modeling for gene editing dynamics. Curr. Opin. Struct. Biol. 2026, 98, 103270
91. C. Pindi†, M. Ahsan†, S. Sinha, and G. Palermo*. Graph Attention Neural Networks Reveal TnsC Filament Assembly in a CRISPR-Associated Transposon. Preprint at BioRxiv. doi: https://doi.org/10.1101/2025.06.17.659969.
90. M. Ahsan, A. Saha, D. Ramos, I. Strohkendl, A. L. Knight, E. Skeens, G. P. Lisi, D. W. Taylor, and G. Palermo*. A Cryptic Binding Pocket Regulates the Metal-Dependent Activity of Cas9. Preprint at BioRxiv. doi: https://doi.org/10.1101/2025.08.25.672025.
2025
89. F. Vieyra†, C. Pindi†, G. P. Lisi, U. N. Morzan* and G. Palermo*. Design Rules for Expanding PAM Compatibility in CRISPR-Cas9 from the VQR, VRER and EQR variants. J. Phys. Chem. B. 2025, 129, 11949–11958. Special Issue “Molecular Simulation and Computational Chemistry: The Legacy of Peter A. Kollman”.
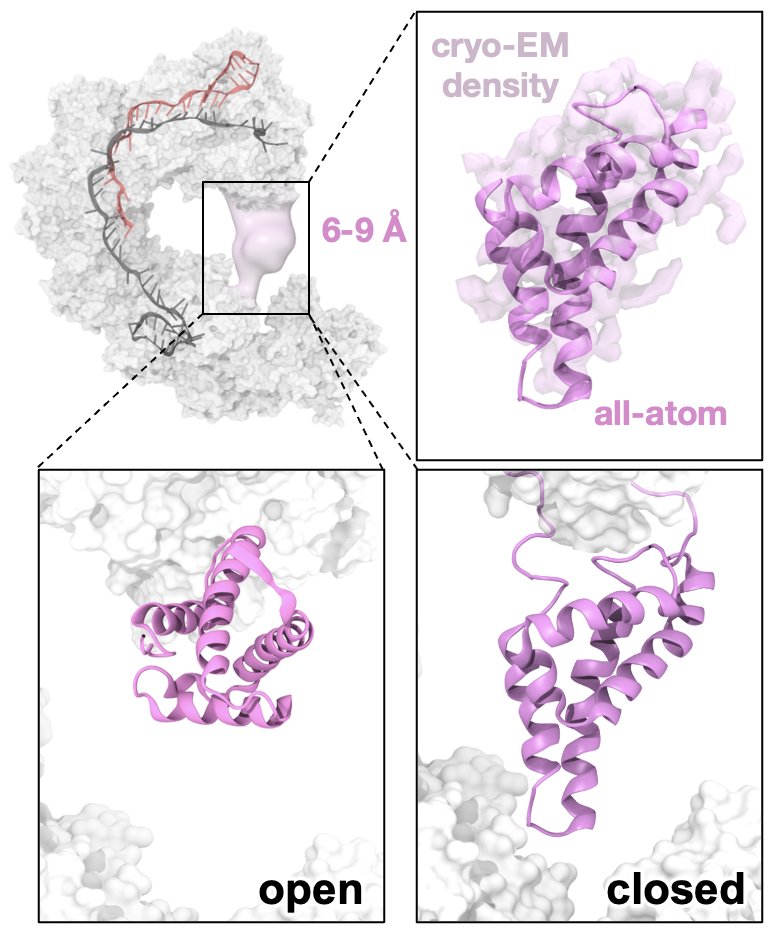
88. A. C. Patel, S. Sinha, P. R. Arantes and G. Palermo*. Unveiling Cas8 Dynamics and Regulation within a transposon-encoded Cascade-TniQ Complex. Proc. Natl. Acad. Sci. USA, 2025, 122, e2422895122.
87. M. Orozco et al. The need to implement FAIR principles in biomolecular simulations. Nat. Methods 2025, 22, 641-645.
86. M. Ahsan†, C. Pindi† and G. Palermo*. Emerging Mechanisms of Metal-Catalyzed RNA and DNA Modifications. Annu. Rev. Phys. Chem. 2025, 76, 497-518.
85. K. A. Hossain, L. Nierzwicki, M. Orozco, J. Czub and G. Palermo*. Flexibility in PAM Recognition Expands DNA Targeting in xCas9. eLife, 2025, 13, RP102538.
84. A. Newman, A. Saha, L. Starrs, P. R. Arantes, G. Palermo, G. Burgio*. CRISPR-Cas12a REC2–Nuc interactions drive target-strand cleavage and constrain trans cleavage. Nucleic Acids Res. 2025, 53, gkaf988.
83. P. R. Arantes, S. Sinha, and G. Palermo*. ParametrizANI: Fast and Accessible Dihedral Parametrization for Small Molecules. J. Chem. Inf. Model. 2025, 65, 11357–11365.
82. H. B. Belato, A. L. Knight, A. M. D’Ordine, C. Pindi, Z. Fan, J. Luo, G. Palermo, G. Jogl and G. P Lisi. Structural and Dynamic Impacts of Single-atom Disruptions to Guide RNA Interactions within the Recognition Lobe of Geobacillus stearothermophilus Cas9. eLife, 2025, 13, RP99275.

2024
81. P. R. Arantes,* X. Chen,* S. Sinha,* A. Saha,* A. C. Patel, M. Sample, L. Nierzwicki, Audrone Lapinaite*, and G. Palermo.* Dimerization of the Deaminase Domain and Locking Interactions with Cas9 Boost Base Editing Efficiency in ABE8e. Nucleic Acids Res, 2024, gkae1066.
80. A. C. Patel, S. Sinha and G. Palermo.* Graph Theory Approaches for Molecular Dynamics Simulations. Q. Rev. Biophys. 2024, 57, e15.
Credits: Dr. Pablo R. Arantes
79. A. Saha, M. Ahsan,† P. R. Arantes,† M. Schmitz, C. Chanez, M. Jinek and G. Palermo.* An Alpha-helical Lid Guides the Target DNA toward Catalysis in CRISPR-Cas12a. Nat. Commun. 2024, 15, 1473. (read PDF).
78. E. Skeens,† S. Sinha,† M. Ahsan, A. M. D’Ordine, G. Jogl, G. Palermo*, and G. P Lisi.* High-Fidelity, Hyper-Accurate, and Evolved Mutants Rewire Atomic Level Communication in CRISPR-Cas9. 2024 Science Adv. 2024, 10, 1045. (read PDF).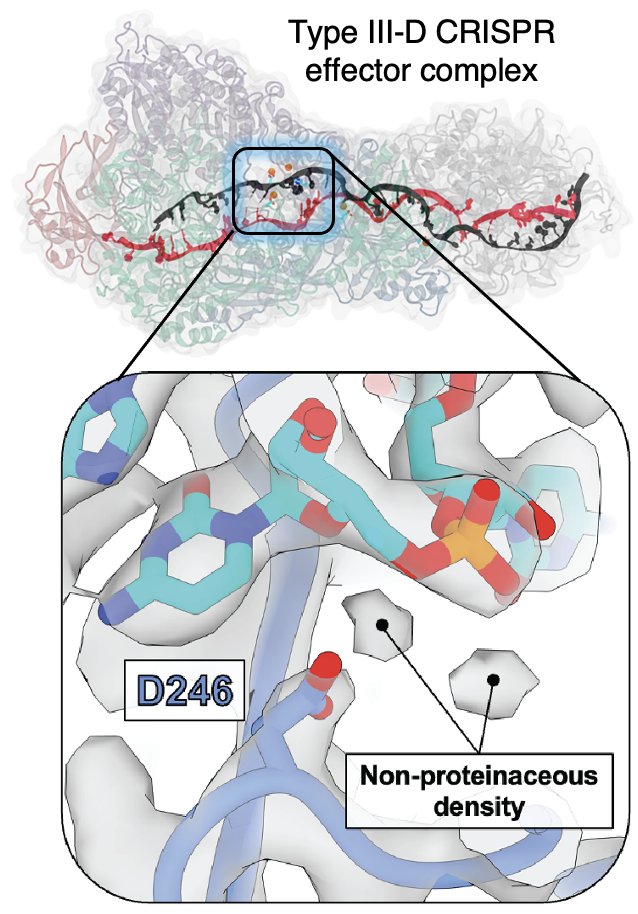
77. E. Schwartz,† J. Bravo,† M. Ahsan,† L. Macias, C. McCafferty, T. Dangerfield, J. Walker, J. Brodbelt, G. Palermo,* P. Fineran, R. Fagerlund† and D. Taylor.† Type III CRISPR-Cas effectors act as protein-assisted ribozymes during RNA cleavage. Nat. Commun. 2024, 15, 3324. (read PDF)
76. I. Strohkendl, A. Saha, C. Moy, A. H. Nguyen, M. Ahsan, R. Russell, G. Palermo, D. W. Taylor.† Cas12a domain flexibility guides R-loop formation and forces RuvC resetting. Mol. Cell. 2024, 84, 1-15. (read PDF).
75. G. Bussi et al. RNA dynamics from experimental and computational approaches. Structure 2024, 32, 1281-1287. (read PDF).
74. X. Fan, Y. Ye, A. Saha, L. Peng, C. Pindi, Q. Wang, L. Yang, J. Liu, X. Tang, G. Palermo, J. Liao, T. Xu, Y. Lu, G. Du. Fine-tuning pH sensor H98 by remote essential residues in the hydrogen-bond network of mTASK-3. Int. J. Biol. Macromol. 2024 13, 132892. (read PDF).
2023
73. G. Palermo* and T. A. Soares.† Editing DNA and RNA through Computations. J. Chem. Inf. Model. 2023, 63, 7603–7604. (read PDF).
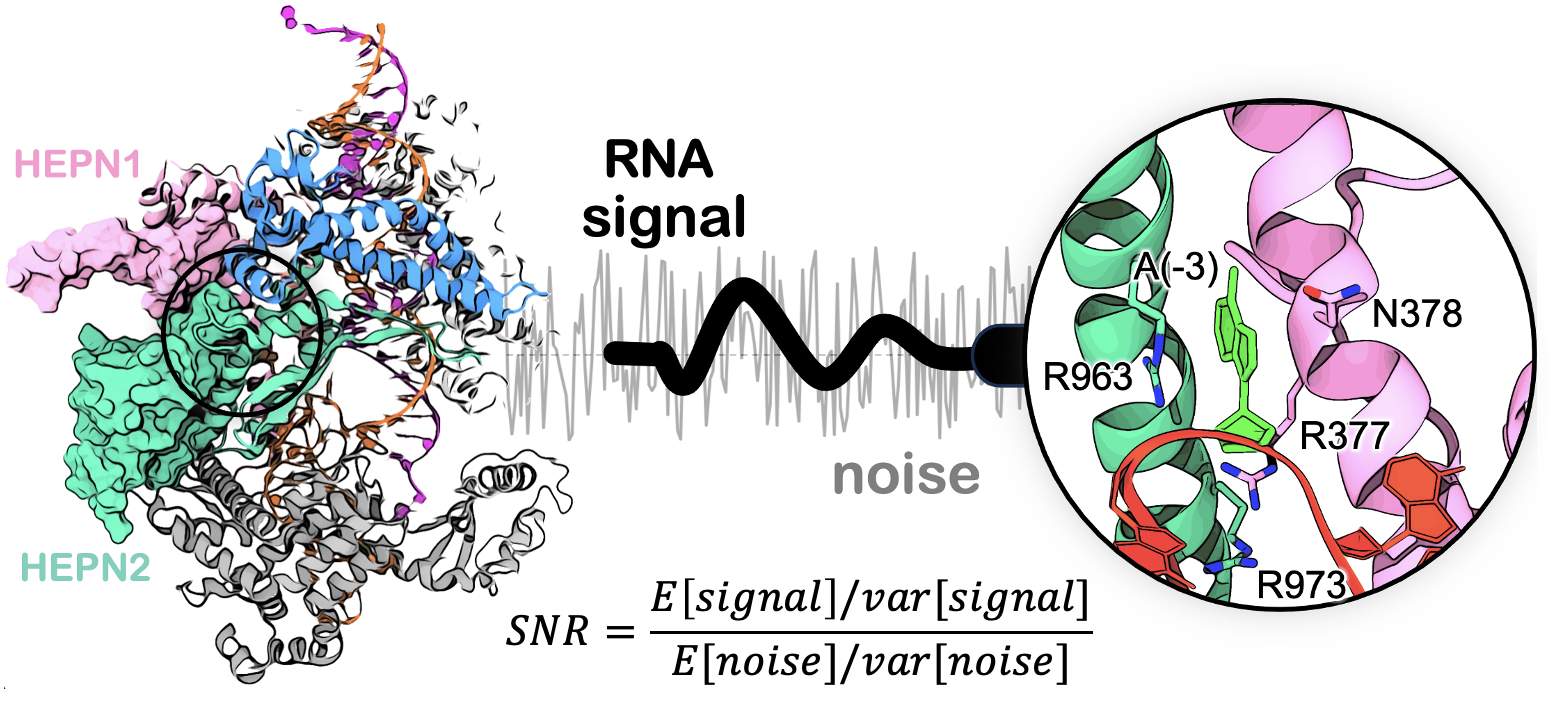
72. S. Sinha, A. M. Molina Vargas, P. R. Arantes,† A. Patel,† M R. O’Connell* and G. Palermo*. Unveiling the RNA-mediated allosteric activation discloses functional hotspots in CRISPR-Cas13a. Nucleic Acids Res. 2023, gkad1127. (read PDF)
71. A. M. Molina Vargas, R. Osborn, S. Sinha, P. R. Arantes, A. Patel, S. Dewhurst, A. Cameron, D. Hardy, G. Palermo* and M R. O’Connell.* New design strategies for ultra-specific CRISPR-Cas13a-based RNA-diagnostic tools with single-nucleotide mismatch sensitivity. Nucleic Acids Res. 2023 gkad1132. (read PDF).
70. A. Saha, C. Pindi and G. Palermo.* The synchronised catalytic dance of CRISPR-Cas9. Nature Catal. 2023, 6, 870-872. (read PDF).
69. S. Sinha, M. Ahsan and G. Palermo.* Establishing the fundamental rules for genetic code expansion. Nature Chem. 2023, 15, 892-893. (read PDF).
68. J. Wang, F. Maschietto, T. Qiu, P. R. Arantes, E. Skeens, G. Palermo,* G. P. Lisi,* V. S. Batista.* Substrate-independent activation pathways of the CRISPR-Cas9 HNH nuclease. Biophys. J. 2023, 122, 4635–4644. (read PDF).
 67. S. Sinha, C. Pindi, M. Ahsan, P. R. Arantes and G. Palermo.* Machines on Genes through the Computational Microscope. J. Chem. Theory Comput. 2023, 19, 1945-1964.
67. S. Sinha, C. Pindi, M. Ahsan, P. R. Arantes and G. Palermo.* Machines on Genes through the Computational Microscope. J. Chem. Theory Comput. 2023, 19, 1945-1964.
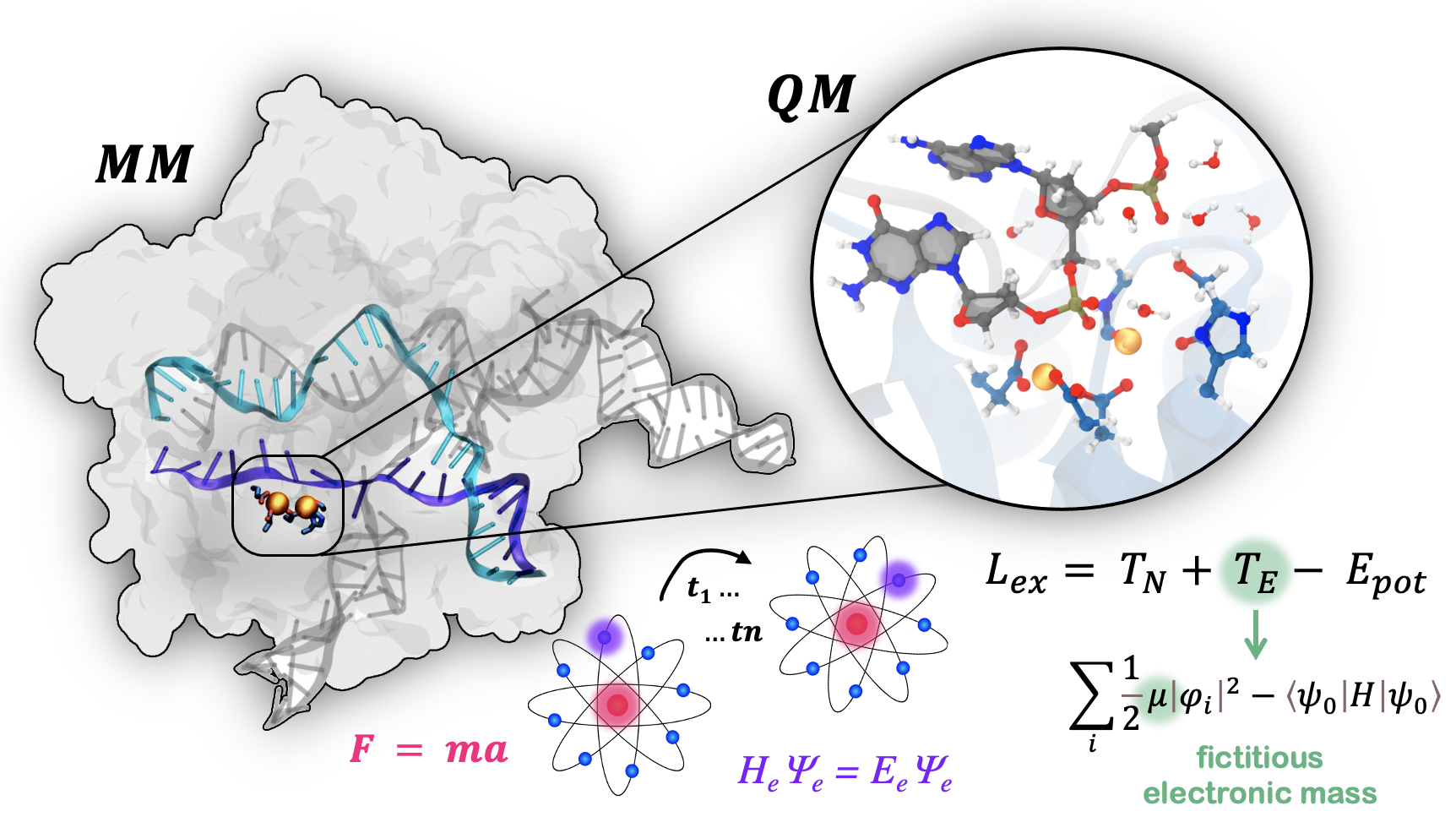
66. Ł. Nierzwicki, M. Ahsan and G. Palermo.* The Electronic Structure of Genome Editors from the First Principles. Electronic Structure, 2023, 5, 014003.
65. A. Kumar, P. R. Arantes, A. Saha, G. Palermo, B. M. Wong. GPU-Enhanced DFTB Metadynamics for Efficiently Predicting Free Energies of Biochemical Systems. Molecules 2023, 28, 1277.
64. J. Wang,† P. R. Arantes, M. Ahsan, S. Sinha, G. W. Kyro, F. Maschietto, B. Allen, E. Skeens, G. P. Lisi,* V. S. Batista† and G. Palermo.* Twisting and Swiveling Domain Motions in Cas9 to Recognize Target DNA Duplex, Make Double-Strand Breaks, and Release Cleaved Duplexes. Front. Mol. Biosci. 2023, 9:1072733.
2022
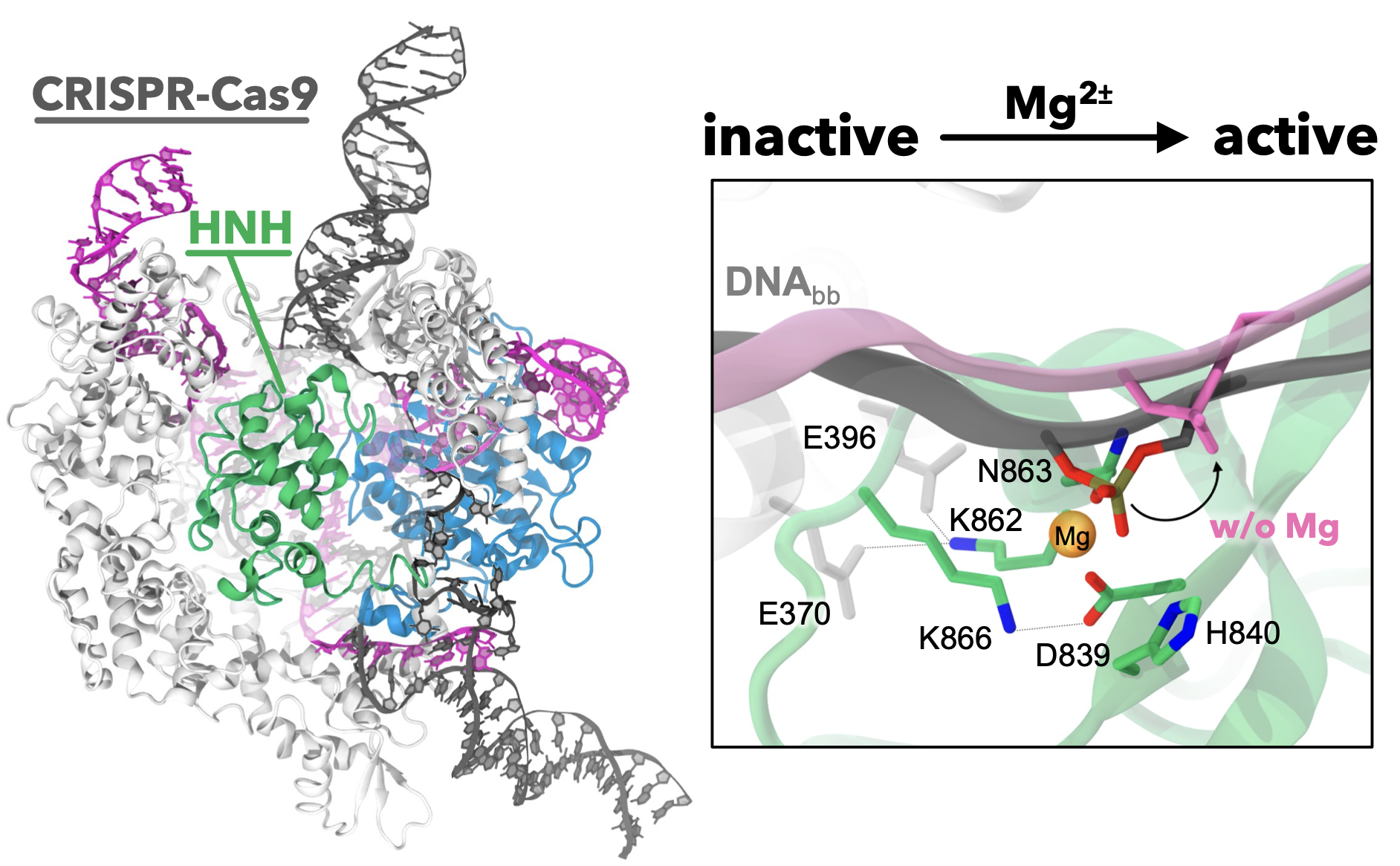 63. Ł. Nierzwicki, K. W. East, J. Binz, R. V. Hsu, M. Ahsan, P. R. Arantes, E. Skeens, M. Pacesa, M. Jinek, G. P. Lisi* and G. Palermo.* Principles of Target DNA Cleavage and Role of Mg2+ in the Catalysis of CRISPR-Cas9. Nature Catal. 2022, 5, 912-922. PDF
63. Ł. Nierzwicki, K. W. East, J. Binz, R. V. Hsu, M. Ahsan, P. R. Arantes, E. Skeens, M. Pacesa, M. Jinek, G. P. Lisi* and G. Palermo.* Principles of Target DNA Cleavage and Role of Mg2+ in the Catalysis of CRISPR-Cas9. Nature Catal. 2022, 5, 912-922. PDF
62. M. Pacesa, C-H. Lin, A. Cléry, A. Saha, P. R. Arantes, K. Bargsten, M. J. Irby, F. H.T. Allain, G. Palermo, P. Cameron, P. D. Donohoue and M. Jinek. Structural basis for Cas9 off-target activity. Cell 2022, 185, 4067-4081.
61. H. B. Belato, C. Norbrun, J. Luo, C. P., S. Sinha, A. M. D’Ordine, G. Jogl, G. Palermo* and G. P. Lisi.* Disruption of Electrostatic Contacts in the HNH Nuclease from a Thermophilic Cas9 Rewires Allosteric Motions and Enhances High-Temperature DNA Cleavage. J. Chem. Phys. 2022, 157, 225103.
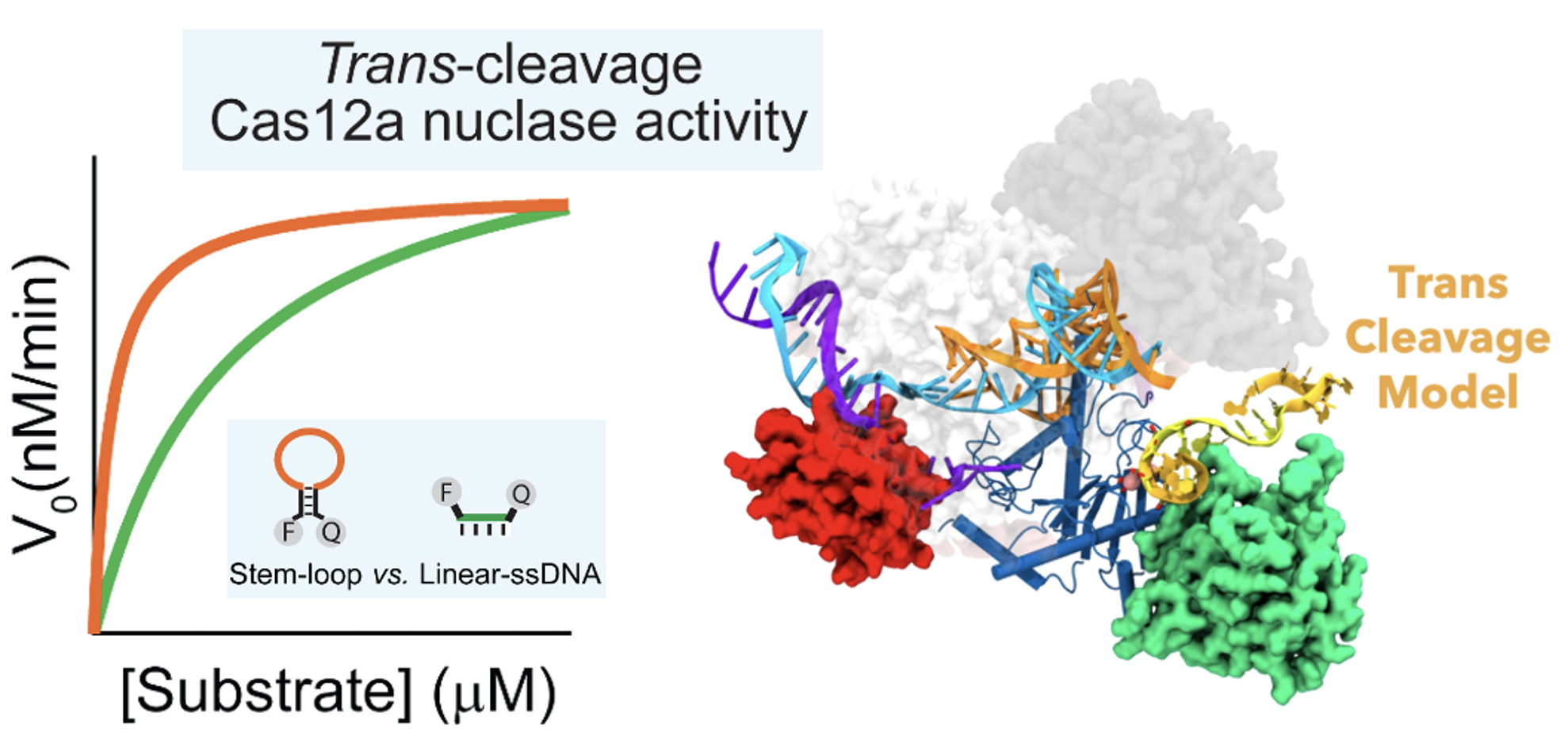 60. M. Rossetti, R. Merlo, N. Bagheri, D. Moscone, A. Valenti, A. Saha, P. R. Arantes, R. Ippodrino, F. Ricci, I. Treglia, E. Delibato, J. van der Oost, G. Palermo.* G. Perugino* and A. Porchetta.* Enhancement of CRISPR/Cas12a trans-cleavage Activity Using Hairpin DNA Reporters. Nucleic Acids Res. 2022, gkac578.
60. M. Rossetti, R. Merlo, N. Bagheri, D. Moscone, A. Valenti, A. Saha, P. R. Arantes, R. Ippodrino, F. Ricci, I. Treglia, E. Delibato, J. van der Oost, G. Palermo.* G. Perugino* and A. Porchetta.* Enhancement of CRISPR/Cas12a trans-cleavage Activity Using Hairpin DNA Reporters. Nucleic Acids Res. 2022, gkac578.
59. G. Palermo.* Celebrating the Next Generation of Women in Computational Chemistry to Increase Diversity, Equity, Inclusion, and Respect. Editorial in J. Chem. Inf. Model. 2022, 62, 3133–3134. JOURNAL COVER ART.
58. A. Saha,† P. R. Arantes† and G. Palermo.* Dynamics and Mechanism of CRISPR-Cas9 through the Lens of Computational Methods. Current Opin. Struct. Biol. 2022, 75, 102400.
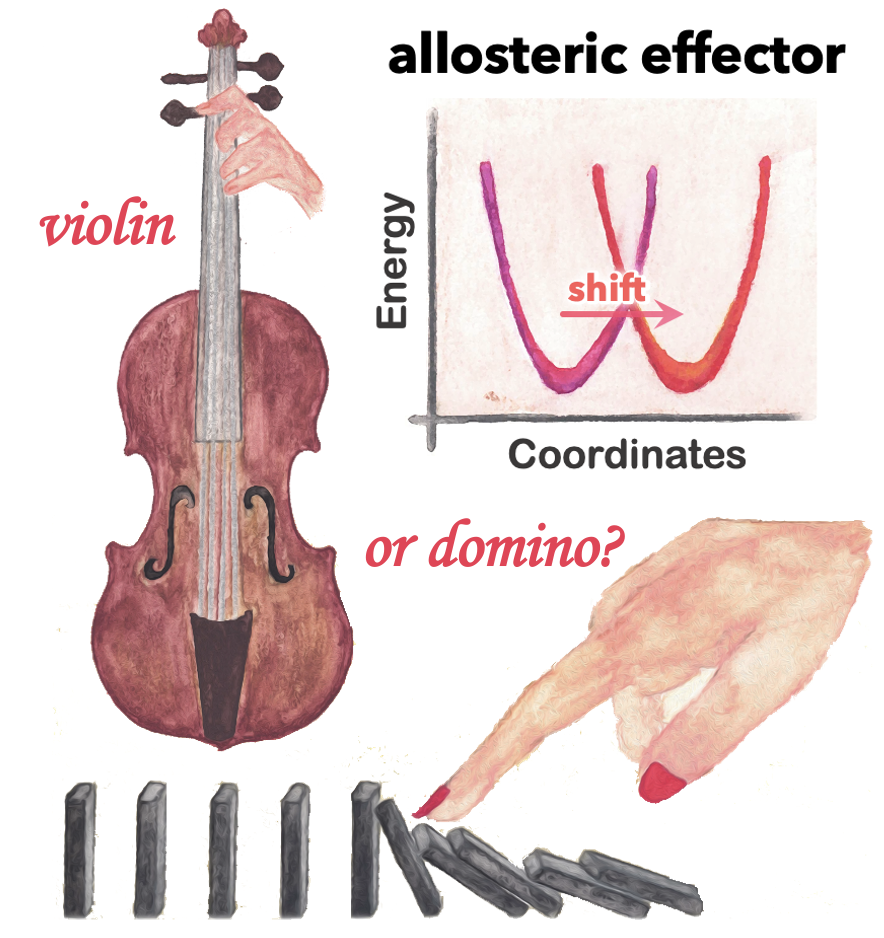 57. Emerging Methods and Applications to Decrypt Allostery in Proteins and Nucleic Acids. P. R. Arantes, A. C. Patel and G. Palermo.* J. Mol. Biol. 2022, 167518.
57. Emerging Methods and Applications to Decrypt Allostery in Proteins and Nucleic Acids. P. R. Arantes, A. C. Patel and G. Palermo.* J. Mol. Biol. 2022, 167518.
56. J. Wang, E. Skeens, P. R. Arantes, F. Maschietto, B. Allen, G. Kyro, G. P. Lisi,* G. Palermo,* V. S. Batista.* Structural Basis for Reduced Dynamics of Three Engineered HNH Endonuclease Lys-to-Ala Mutants of the Clustered Regularly Interspaced Short Palindromic Repeat-Associated 9 Enzyme. Biochemistry 2022 61, 785–794.
55. Structural and Dynamic Insights into the HNH Nuclease of Divergent Cas9 Species. H. B. Belato, A. M. D’Ordine, Ł. Nierzwicki, P. R. Arantes, G. Jogl, G. Palermo* and G. P. Lisi.* J. Struct. Biol. 2022, 214, 107814.
2021
 54. Enhanced Specificity Mutations Perturb Allosteric Signaling in CRISPR-Cas9. Ł. Nierzwicki, K. W. East, U. N. Morzan, P. R. Arantes, V. S. Batista, G. P. Lisi* and G. Palermo.* eLife 2021, 10: e73601.JOURNAL COVER ART.
54. Enhanced Specificity Mutations Perturb Allosteric Signaling in CRISPR-Cas9. Ł. Nierzwicki, K. W. East, U. N. Morzan, P. R. Arantes, V. S. Batista, G. P. Lisi* and G. Palermo.* eLife 2021, 10: e73601.JOURNAL COVER ART.
53. J. Wang,† P. R. Arantes,† A. Bhattarai, R. V. Hsu, S. Pawnikar, Y. M. Huang, G. Palermo* and Y. Miao.* Gaussian accelerated molecular dynamics (GaMD): principles and applications. WIREs Comp. Mol. Sci. 2021, e1521.
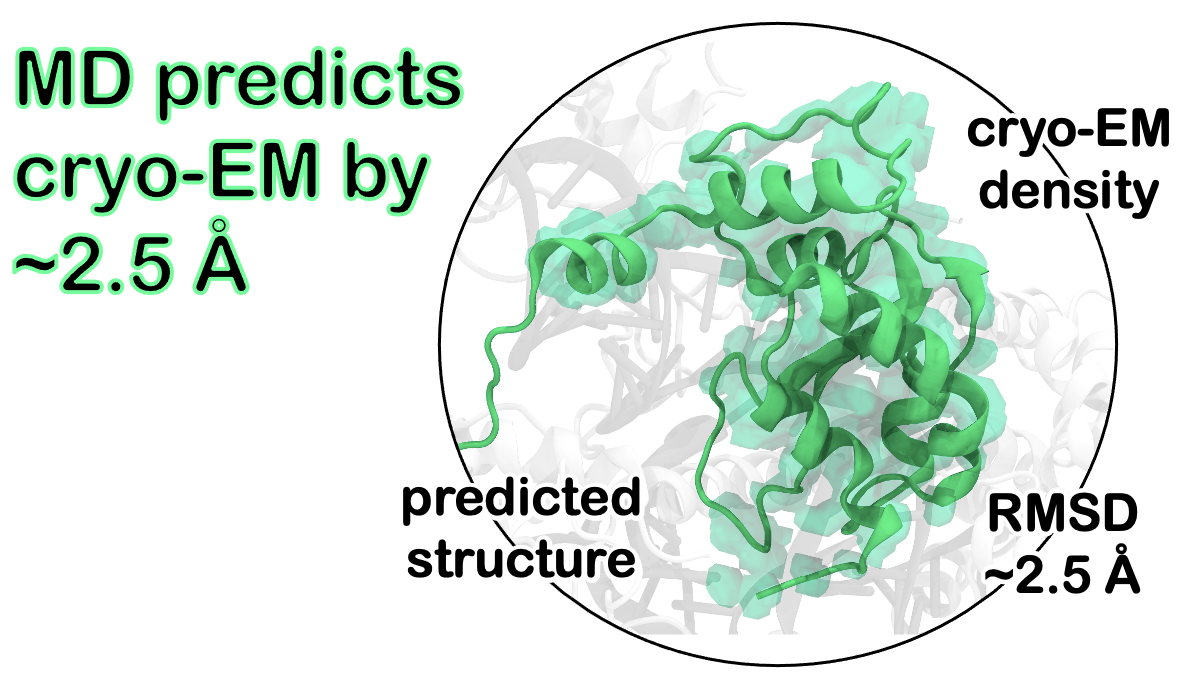 52. Ł. Nierzwicki and G. Palermo.* Molecular Dynamics to predict Cryo-EM: capturing transitions and short-lived conformational states of biomolecules. Front. Mol. Biosci. 2021, 8:641208.
52. Ł. Nierzwicki and G. Palermo.* Molecular Dynamics to predict Cryo-EM: capturing transitions and short-lived conformational states of biomolecules. Front. Mol. Biosci. 2021, 8:641208.
 51. J. Manigrasso, M. De Vivo* and G. Palermo.* Controlled Trafficking of Multiple Cations Prompt Nucleic Acid Hydrolysis. ACS Catalysis 2021, 11, 14, 8786–8797. JOURNAL COVER ART.
51. J. Manigrasso, M. De Vivo* and G. Palermo.* Controlled Trafficking of Multiple Cations Prompt Nucleic Acid Hydrolysis. ACS Catalysis 2021, 11, 14, 8786–8797. JOURNAL COVER ART.
50. N. T. Zewde, R. V. Hsu. D. Morikis* and G. Palermo.* Systems Biology Modeling of the Complement System Under Immune Susceptible Pathogens. Front. Phys. 2021, 9, 603704.
49. N. Farag, R. Mattossovich, R. Merlo, Ł. Nierzwicki, G. Palermo, A. Porchetta,† G. Perugino† and F. Ricci.† Folding-upon-repair DNA nanoswitches for monitoring DNA repair enzymes activity. Angew. Chem. Int. Ed. 2021, 133, 2-9.
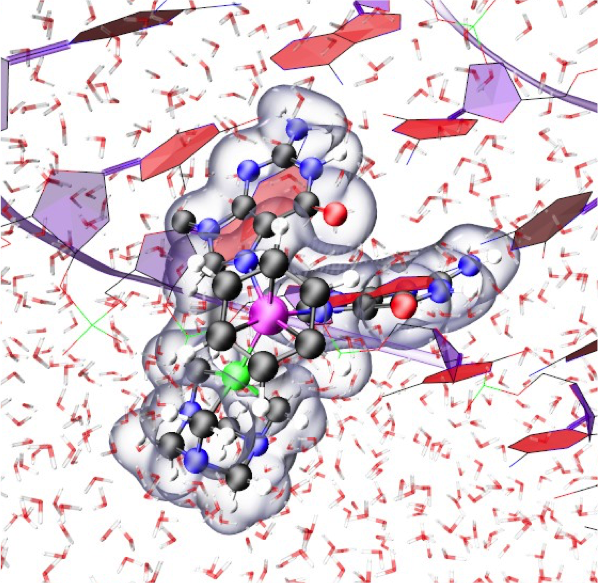 48. G. Palermo, A. Spinello, A. Saha, and A. Magistrato. Frontiers of Metal-coordinating Drug Design. Expert Opin. Drug. Disc. 2021, 1-15.
48. G. Palermo, A. Spinello, A. Saha, and A. Magistrato. Frontiers of Metal-coordinating Drug Design. Expert Opin. Drug. Disc. 2021, 1-15.
47. Y. B. Narkhede,† A. K. Gautam,† R. V. Hsu,† W. Rodriguez, N. T. Zewde, R. E. S. Harrison, P. R. Arantes, Z. Gaieb, R. D. Gorham Jr, C. Kieslich, D. Morikis, A. Sahu* and G. Palermo.* Role of Electrostatic Hotspots in the Selectivity of Complement Control Proteins toward Human and Bovine Complement Inhibition. Front. Mol. Biosci. 2021, 8:618068.
2020
 46. K. W. East, J. C. Newton, U. N. Morzan, Y. B. Narkhede, A. Acharya, E. Skeens, G. Jogl, V. S. Batista, G. Palermo* and G. P. Lisi.* Allosteric Motions of the CRISPR-Cas9 HNH Nuclease Probed by NMR and Molecular Dynamics. J. Am. Chem. Soc. 2020, 142, 1348-1358.
46. K. W. East, J. C. Newton, U. N. Morzan, Y. B. Narkhede, A. Acharya, E. Skeens, G. Jogl, V. S. Batista, G. Palermo* and G. P. Lisi.* Allosteric Motions of the CRISPR-Cas9 HNH Nuclease Probed by NMR and Molecular Dynamics. J. Am. Chem. Soc. 2020, 142, 1348-1358.
Featured in the JACS 2021 Early Investigators Virtual Issue!
 45. L. Casalino, Ł. Nierzwicki, M. Jinek and G. Palermo.* Catalytic Mechanism of Non-Target DNA Cleavage in CRISPR-Cas9 Revealed by Ab-Initio Molecular Dynamics. ACS Catalysis, 2020, 22, 13596-13605. JOURNAL COVER ART.
45. L. Casalino, Ł. Nierzwicki, M. Jinek and G. Palermo.* Catalytic Mechanism of Non-Target DNA Cleavage in CRISPR-Cas9 Revealed by Ab-Initio Molecular Dynamics. ACS Catalysis, 2020, 22, 13596-13605. JOURNAL COVER ART.
44. B. P. Mitchell, R. V. Hsu, M. A. Medrano, N. T. Zewede, Y. B. Narkhede, G. Palermo.* Spontaneous embedding of DNA mismatches within the RNA:DNA hybrid in CRISPR-Cas9. Front. Mol. Biosci. 2020, 7, 39.
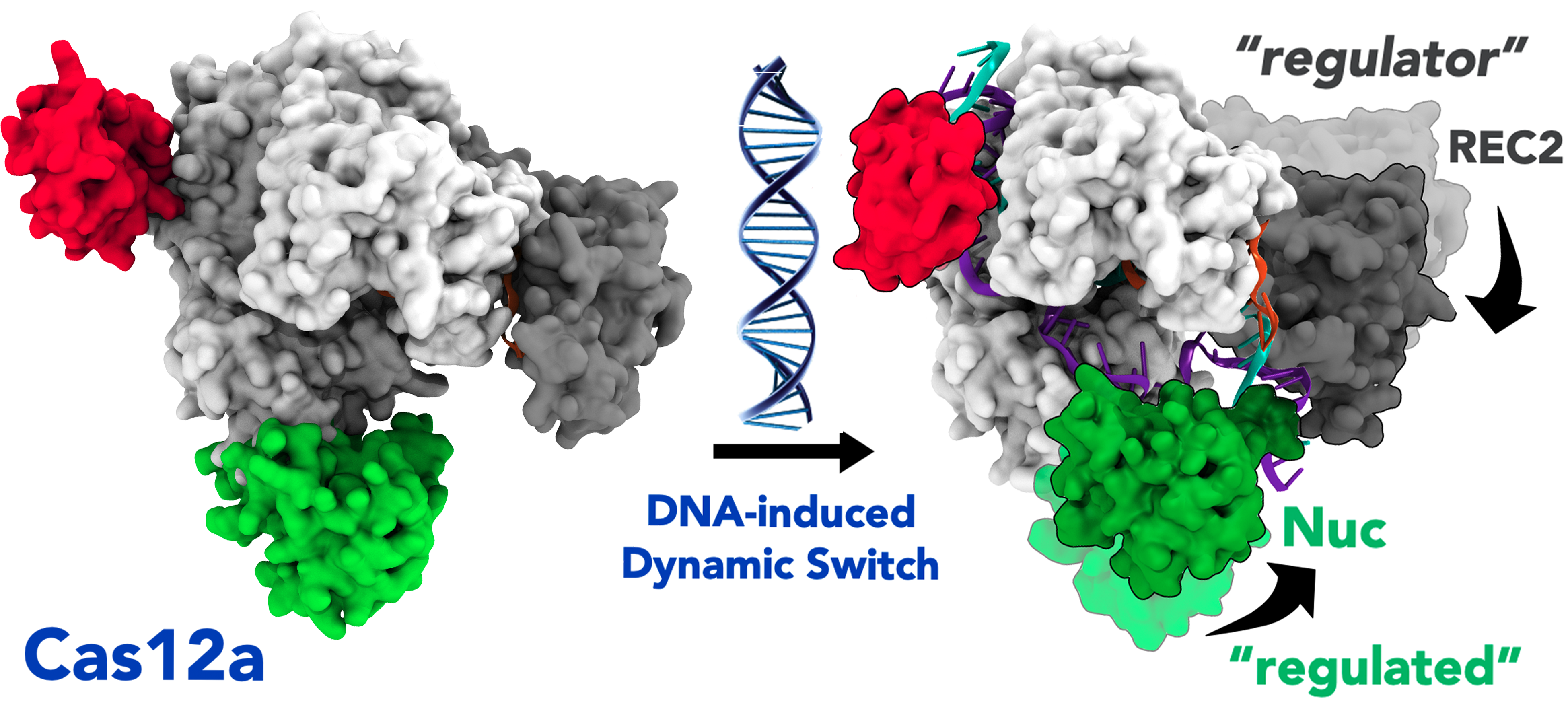 43. A. Saha,† P. R. Arantes,† R. V. Hsu, Y. B. Narkhede, M. Jinek and G. Palermo.* Molecular Dynamics Reveal a DNA-Induced Dynamic Switch Triggering Activation of CRISPR-Cas12a. J. Chem. Inf. Model. 2020, 60, 6427–6437. JOURNAL COVER ART.
43. A. Saha,† P. R. Arantes,† R. V. Hsu, Y. B. Narkhede, M. Jinek and G. Palermo.* Molecular Dynamics Reveal a DNA-Induced Dynamic Switch Triggering Activation of CRISPR-Cas12a. J. Chem. Inf. Model. 2020, 60, 6427–6437. JOURNAL COVER ART.
ACS Editors’ Choice.
 42. Ł. Nierzwicki, P. R. Arantes, A. Saha and G. Palermo.* Establishing the Allosteric Mechanism in CRISPR-Cas9. WIREs Comp. Mol. Biosci. 2020, e1503. JOURNAL COVER ART.
42. Ł. Nierzwicki, P. R. Arantes, A. Saha and G. Palermo.* Establishing the Allosteric Mechanism in CRISPR-Cas9. WIREs Comp. Mol. Biosci. 2020, e1503. JOURNAL COVER ART.
41. P. R. Arantes, A. Saha and G. Palermo.* Fighting COVID-19 Using Molecular Dynamics Simulations. First Reaction, ACS Cent. Sci. 2020, 6, 1654–1656.
40. R. E. H. Harrison, N. T. Zewde, Y. B. Narkhede, R. V. Hsu, D. Morikis, V. Vullev and G. Palermo.* Factor-H Inspired Design of Peptide Biomarkers of the Complement C3d Protein. ACS Med. Chem. Lett. 2020, 11, 1054-1059.
39. J. Cui, F. Zhang, Ł. Nierzwicki, G. Palermo, R. Linhardt and G. P. Lisi. Mapping the Structural and Dynamic Determinants of pH-sensitive Heparin Binding to Granulocyte Macrophage-colony Stimulating Factor. Biochemistry 2020, 59, 3541–3553.
38. Skeens, J. Y. Cui, H. B. Belato, B. P. Mitchell, R. V. Hsu, V. S. Batista, G. Palermo, G. P. Lisi. NMR and computational methods for molecular resolution of allosteric pathways in enzyme complexes. Biophys. Rev. 2020, 1-20 .
37. G. Palermo, Y. Sugita, W. Wriggers and R. E. Amaro. Faces of Contemporary CryoEM Information and Modeling. J. Chem. inf. Model. 2020, 60, 2407-2409.
 36. A. Palermo, L. Casalino & G. Palermo.* Vincent Van Gogh’ Autocatalysis. The Art of Theoretical Biology, Springer Nature 2020, 114-115.
36. A. Palermo, L. Casalino & G. Palermo.* Vincent Van Gogh’ Autocatalysis. The Art of Theoretical Biology, Springer Nature 2020, 114-115.
35. G. Palermo, A. M.J.J. Bonvin, M. Dal Peraro, R. E Amaro and V. Tozzini. Multiscale Modeling from Macromolecules to Cell: Opportunities and Challenges of Biomolecular Simulations. Front. Mol. Biosci. 2020, 7, 194.
2019

34. S. Vanni, L. Riccardi, G. Palermo and M. De Vivo. Structure and dynamics of the acyl chains in the membrane trafficking and enzymatic processing of lipids. Acc. Chem. Res. 2019, 52, 3087-3096. JOURNAL COVER ART.
33. G. Palermo.* Dissecting Structure and Function of RNA:DNA Hybrids. Chem 2019, 5, 1364-1366.
 32. G. Palermo.* Structure and Dynamics of the CRISPR-Cas9 Catalytic Complex. J. Chem. Inf. Model. 2019, 59, 2394-2406. JOURNAL COVER ART.
32. G. Palermo.* Structure and Dynamics of the CRISPR-Cas9 Catalytic Complex. J. Chem. Inf. Model. 2019, 59, 2394-2406. JOURNAL COVER ART.
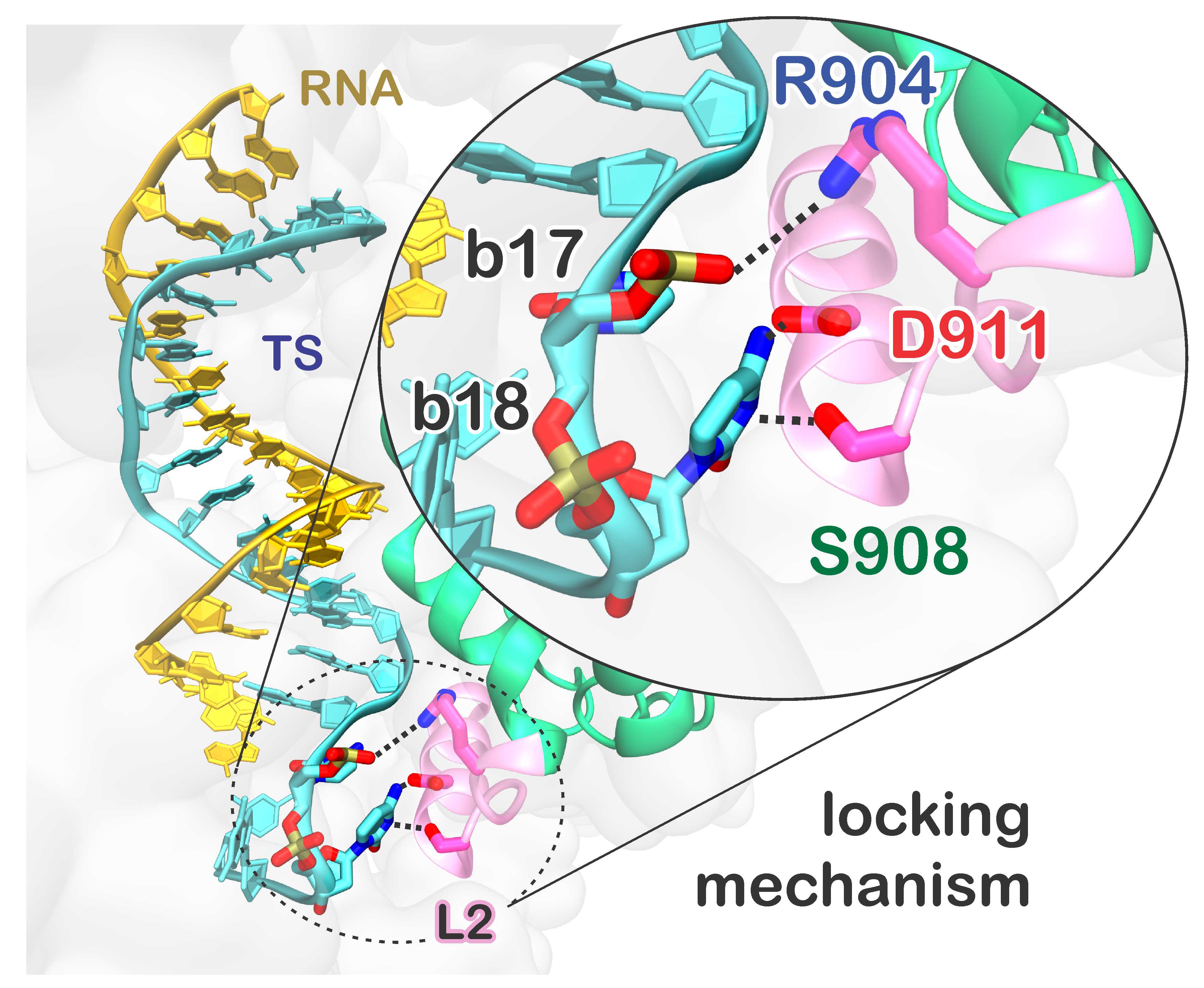
31. C. G. Ricci, J. S. Chen, Y. Miao, M. Jinek, J. A. Doudna, J. A. McCammon and G. Palermo.* Deciphering Off-target Effects in CRISPR-Cas9 through Accelerated Molecular Dynamics. ACS Cent. Sci. 2019, 5, 651-662.
30. J. Borisek, A. Saltalamacchia, A. Galli, G. Palermo, E. Molteni, L. Malcovati, A. Magistrato. Disclosing the Impact of Carcinogenic SF3b Mutations on Pre-mRNA Recognition Via All-Atom Simulations. Biomolecules 2019, 9, 633.
 29. G. Palermo,* L. Casalino, A. Magistrato and J. A. McCammon. Understanding the mechanistic basis of non-coding RNA through molecular dynamics simulations. J. Struct. Biol. 2019, 206, 267-279. JOURNAL COVER ART.
29. G. Palermo,* L. Casalino, A. Magistrato and J. A. McCammon. Understanding the mechanistic basis of non-coding RNA through molecular dynamics simulations. J. Struct. Biol. 2019, 206, 267-279. JOURNAL COVER ART.
Cover picture awarded First Place Prize at the 2021 RNA Society Arts and Music competition (drawing category)!
Cover picture awarded First Place Prize at the 2017 Biophysical Society Image Contest!
28. G. Palermo,* C. G. Ricci and J. A. McCammon. The invisible dance of CRISPR-Cas9. Molecular simulations unveil the molecular side of the gene-editing revolution. Physics Today, 2019, 72, 4, 30.
27. S. J. Wodak. et Al. Allostery in its many disguises: from theory to applications. Structure 2019, 4, 566-578.
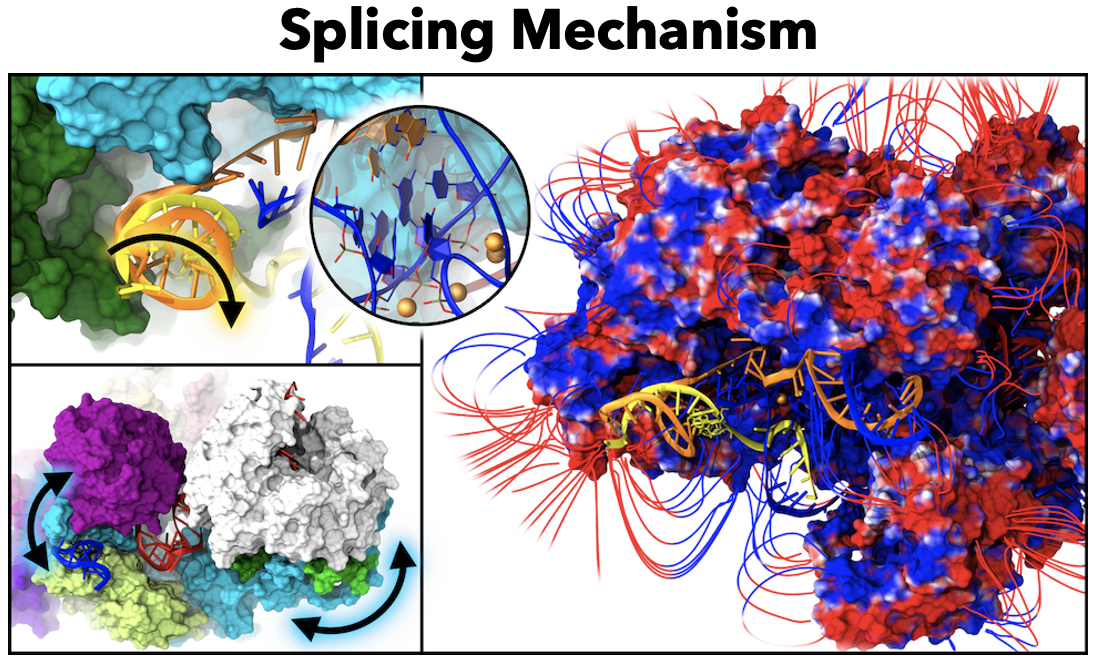 26. L. Casalino, G. Palermo, A. Spinello, U. Rothlisberger and A. Magistrato. All-Atom Simulations Disentangle the Functional Dynamics Underlying Gene Maturation in the Intron Lariat Spliceosome. Proc. Natl. Acad. Sci. USA 2018, 115, 6584-6589.
26. L. Casalino, G. Palermo, A. Spinello, U. Rothlisberger and A. Magistrato. All-Atom Simulations Disentangle the Functional Dynamics Underlying Gene Maturation in the Intron Lariat Spliceosome. Proc. Natl. Acad. Sci. USA 2018, 115, 6584-6589.
25. G. Palermo,* J. S. Chen, C. G. Ricci, I. Rivalta, M. Jinek, V. S. Batista, J. A. Doudna and J. A. McCammon. Key role of the REC lobe during CRISPR-Cas9 activation by “sensing”, “regulating” and “locking” the catalytic HNH domain. Quarterly Rev. Biophys. 2018, 51, e9.
 24. G. Palermo,* K. A. Armacost and M. Nagan. Women Make COMP: Mentoring the Next-Generation of Women in Computational Chemistry. Viewpoint in J. Chem. Inf. Model. 2019, 59, 4061-4062.
24. G. Palermo,* K. A. Armacost and M. Nagan. Women Make COMP: Mentoring the Next-Generation of Women in Computational Chemistry. Viewpoint in J. Chem. Inf. Model. 2019, 59, 4061-4062.
23. G. Palermo, Y. Sugita, W. Wriggers and R. E. Amaro. Frontiers in CryoEM Modeling. J. Chem. Inf. Model. 2019, 59, 3091-3093.
Before UCR
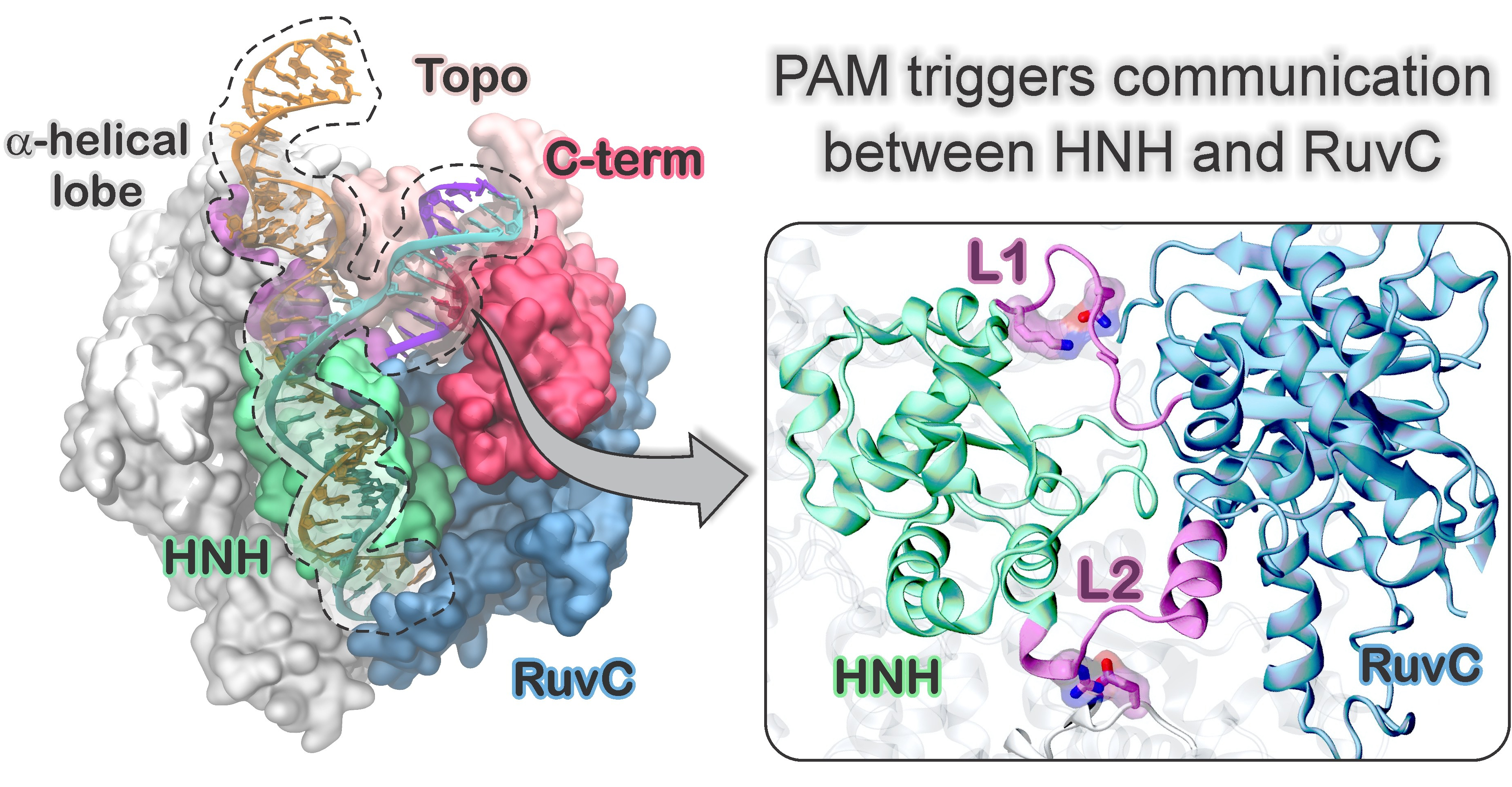 22. G. Palermo,* C. G. Ricci, A. Fernando, R. Basak, M. Jinek, I. Rivalta, V. S. Batista and J. A. McCammon PAM-induced allostery activates CRISPR-Cas9. J. Am. Chem. Soc. 2017, 139, 16028–16031. JOURNAL COVER ART.
22. G. Palermo,* C. G. Ricci, A. Fernando, R. Basak, M. Jinek, I. Rivalta, V. S. Batista and J. A. McCammon PAM-induced allostery activates CRISPR-Cas9. J. Am. Chem. Soc. 2017, 139, 16028–16031. JOURNAL COVER ART.
Featured in the JACS 2018 Early Investigators Virtual Issue!
21. G. Palermo,* Y. Miao, R. C. Walker, M. Jinek and J. A. McCammon.* CRISPR-Cas9 conformational activation as elucidated from enhanced molecular simulations. Proc. Natl. Acad. Sci. USA. 2017, 114, 7260–7265.
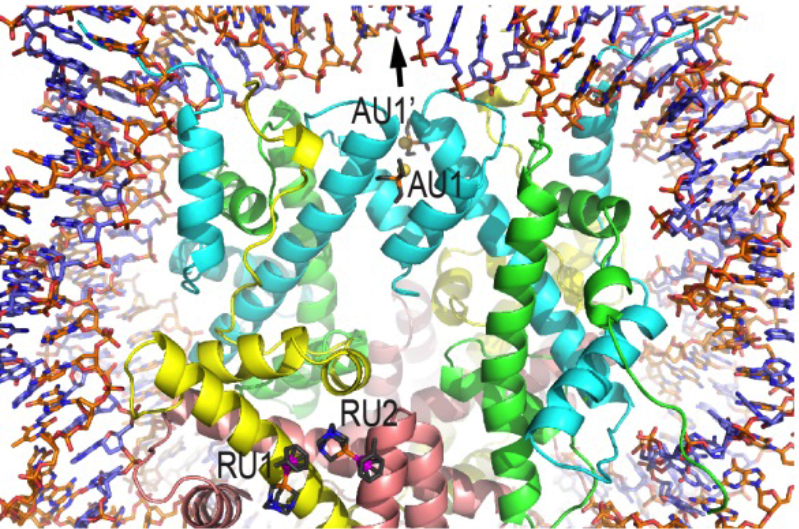
20. Z. Adhireksan,† G. Palermo,† T. Riedel,† Z. Ma, R. Muhammad, P. J. Dyson, U. Rothlisberger and C. A. Davey. Allosteric cross-talk in chromatin can mediate drug-drug synergy. Nat. Commun. 2017, 8, 14860.
 19. N. Ashari Astani, S. Meloni, A. H. Salavati, G. Palermo, M. Graetzel, U. Rothlisberger. Computational Characterization of the Dependence of Halide Perovskite Effective Masses on Chemical Composition and Structure. J. Phys. Chem. C. 2017, 121, 23886–23895.
19. N. Ashari Astani, S. Meloni, A. H. Salavati, G. Palermo, M. Graetzel, U. Rothlisberger. Computational Characterization of the Dependence of Halide Perovskite Effective Masses on Chemical Composition and Structure. J. Phys. Chem. C. 2017, 121, 23886–23895.
18. L. Casalino, G. Palermo, N. Abdurakhmonova, U. Rothlisberger and A. Magistrato. Development of Site-specific Mg-RNA Force Field Parameters: A Dream or Reality? Guidelines from Combined Molecular Dynamics and Quantum Mechanics Simulations. J. Chem. Theory Comput. 2017, 13, 340–352.
 17. G. Palermo,* Y. Miao, R. C. Walker, M. Jinek and J. A. McCammon. Striking plasticity of CRISPR-Cas9 and key role of non-target DNA, as revealed from molecular simulations. ACS Cent. Sci. 2016, 2, 756–763.
17. G. Palermo,* Y. Miao, R. C. Walker, M. Jinek and J. A. McCammon. Striking plasticity of CRISPR-Cas9 and key role of non-target DNA, as revealed from molecular simulations. ACS Cent. Sci. 2016, 2, 756–763.
First computer simulations of CRISPR-Cas9.
Awarded the 2017 Best Use of HPC in Life Sciences.
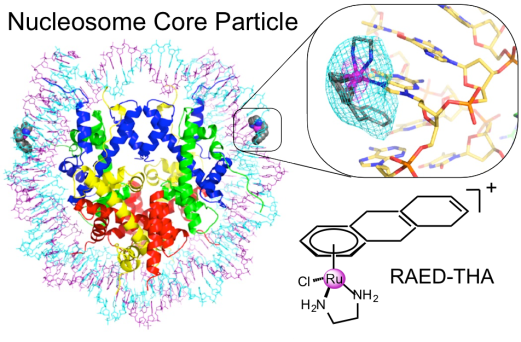
16. Z. Ma,† G. Palermo,† Z. Adhireksan,† B. S. Murray, T. von Erlach, P. J. Dyson, U. Rothlisberger and C. A. Davey. An Organometallic Compound Displays a Unique One-Stranded Intercalation Mode that is DNA Topology-Dependent. Angew. Chem. Int. Ed. 2016, 128, 7441–7444.

15. L. Casalino, G. Palermo, U. Rothlisberger and A. Magistrato. Who Activates the Nucleophile in Ribozyme Catalysis? An Answer from the Splicing Mechanism of Group II Introns. J. Am. Chem. Soc. 2016, 138, 10374–10377. JOURNAL COVER ART.
14. G. Palermo, A. Magistrato, T. Riedel, T. von Erlach, C. A. Davey, P. J. Dyson and U. Rothlisberger. Fighting cancer with organometallic compounds: from naked DNA to protein and chromatin targeting strategies. ChemMedChem, 2016, 11, 1199–1210. Special Issue on Polypharmacology and Multitarget Drugs.
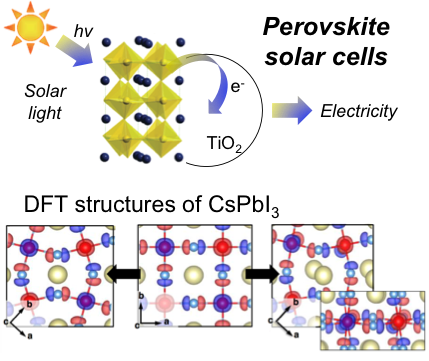 13. S. Meloni, G. Palermo, N. Ashari Astani, M. Graetzel and U. Rothlisberger. Valence and conduction bands engineering in halide perovskites for solar cell applications. J. Mater. Chem. A. 2016, 4, 15997–16002.
13. S. Meloni, G. Palermo, N. Ashari Astani, M. Graetzel and U. Rothlisberger. Valence and conduction bands engineering in halide perovskites for solar cell applications. J. Mater. Chem. A. 2016, 4, 15997–16002.
12. G. Palermo, A. D. Favia, M. Convertino and M. De Vivo. The molecular basis for dual Fatty Acid Amide Hydrolase (FAAH)/Cyclooxygenase (COX) inhibition. ChemMedChem, 2016, 11, 1252–1258.

11. G. Palermo, A. Cavalli, M. L. Klein, M. Alfonso Prieto, M. Dal Peraro and M. De Vivo. Catalytic metal ions and enzymatic processing of DNA and RNA. Acc. Chem. Res. 2015, 48, 220–228.

10. G. Palermo, I. Bauer, P. Campomanes, A. Cavalli, A. Armirotti, S. Girotto, U. Rothlisberger and M. De Vivo. Keys to lipid selection in FAAH catalysis: structural flexibility, gating residues, and multiple binding pockets. PLoS Comput. Biol. 2015, 11, e1004231. JOURNAL COVER ART.
9. G. Palermo, P. Campomanes, A. Cavalli, U. Rothlisberger and M. De Vivo. Anandamide hydrolysis in FAAH reveals a dual strategy for efficient enzyme-assisted amide bond cleavage via nitrogen inversion. J. Phys. Chem. B. 2015, 119, 789–801. Prof. W. Jorgensen Festschrift. JOURNAL COVER ART.
8. G. Palermo, A. Cavalli, U. Rothlisberger and M. De Vivo. Computational insights into function and inhibition of the Fatty Acid Amide Hydrolase (FAAH). Eur. J. Med. Chem. 2015, 91, 15–26.
 7. G. Palermo, E. Minniti, M. L. Greco, L. Riccardi, E. Simoni, M. Convertino, C. Marchetti, M. Rosini, C. Sissi, A. Minarini and M. De Vivo. An optimized polyamine moiety boosts potency of human type II topoisomerase poisons as quantified by comparative analysis centered on the clinical candidate F14152. Chem. Commun. 2015, 51, 14310
7. G. Palermo, E. Minniti, M. L. Greco, L. Riccardi, E. Simoni, M. Convertino, C. Marchetti, M. Rosini, C. Sissi, A. Minarini and M. De Vivo. An optimized polyamine moiety boosts potency of human type II topoisomerase poisons as quantified by comparative analysis centered on the clinical candidate F14152. Chem. Commun. 2015, 51, 14310
6. G. Palermo and M. De Vivo. Computational Methods for Drug Discovery. Book chapter in Encyclopedia of Nanotechnology, 2nd Ed. Springer, Heidelberg, 2015, 1-15.
5. G. Palermo, P. Campomanes, M. Neri, D. Piomelli, A. Cavalli, U. Rothlisberger and M. De Vivo. Wagging the tail: essential role of substrate flexibility in FAAH catalysis. J. Chem. Theory. Comput. 2013, 15, 1202–1213.
 4. G. Palermo, M. Stenta, A. Cavalli, M. Dal Peraro and M. De Vivo. Molecular simulations highlight the role of metals in catalysis and inhibition of type II topoisomerase. J. Chem. Theory. Comput. 2013, 9, 857–862. JOURNAL COVER ART.
4. G. Palermo, M. Stenta, A. Cavalli, M. Dal Peraro and M. De Vivo. Molecular simulations highlight the role of metals in catalysis and inhibition of type II topoisomerase. J. Chem. Theory. Comput. 2013, 9, 857–862. JOURNAL COVER ART.
3. G. Palermo, D. Branduardi, M. Masetti, A. Lodola, M. Mor, D. Piomelli, A. Cavalli and M. De Vivo. Covalent inhibitors of fatty acid amide hydrolase: a rationale for the activity of piperidine and piperazine aryl ureas. J. Med. Chem. 2011, 54, 6612–6623.
2. E. Brunk, N. Ashari, P. Athri, P. Campomanes, F. F. de Carvalho, B. F. Curchod, P. Diamantis, M. Doemer, J. Garrec, A. Laktionov, M. Micciarelli, M. Neri, G. Palermo, T. J. Penfold, S. Vanni, I. Tavernelli and U. Rothlisberger. Pushing frontiers of first-principles based computer simulations of chemical and biological systems. Chimia (Aarau), 2011, 65, 667–671.
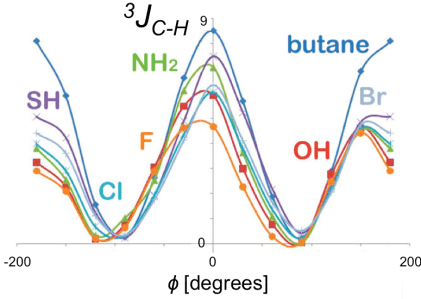
1. G. Palermo, R. Riccio and G. Bifulco. Effect of Electronegative Substituents and Angular Dependence on the Heteronuclear Spin-Spin Coupling Constant 3JC-H: An Empirical Prediction Equation Derived by Density Functional Theory Calculations. J. Org. Chem. 2010, 7, 1982–1991.