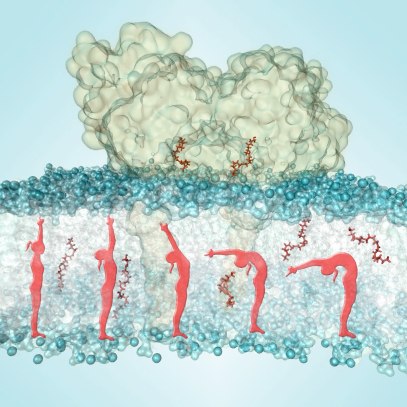
The fatty acid amide hydrolase (FAAH) is a key membrane protein of the endocannabinoid system, which is fundamental for human health and crucial in the regulation of such as pain and inflammation. By acting as an hydrolytic enzyme of the endocannaminoid lipids, FAAH exerts a modulatory action on a variety of inflammatory-related diseases, including cancer. The understanding of its function and inhibition is of crucial importance in the search of novel drugs and therapeutic strategies.
During my graduate studies, I have deciphered the function and inhibition of FAAH via classical & hybrid quantum mechanics/molecular mechanics (QM/MM) approaches. This project has made massive use of the Car-Parrinello approach and of free energy methods decrypting the enzymatic catalysis.
Long timescale simulations of FAAH in complex with its main lipid substrate anandamide in a realistic membrane highlighted the structural flexibility as a key determinant for the substrate recognition and catalysis (Palermo et al. JCTC 2013). By applying Bayesian statistics and an in-house Machine Learning approach, we identified a solid model for the reactant state, clarifying the mechanism of lipid binding for catalysis.
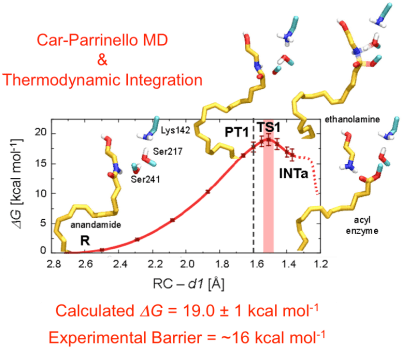
Subsequently, Car-Parrinello QM/MM MD & thermodynamic integration characterized the entire enzymatic cycle, depicting a highly concerted catalysis, which underlies FAAH efficiency in hydrolyzing lipids (Palermo et al. J. Phys. Chem. B. 2015). Car-Parrinello shows that FAAH uses an exquisite catalytic strategy for the amide bond hydrolysis, which can be of general applicability for other amidases/proteases, providing key insights for de novo enzyme design and drug discovery efforts.
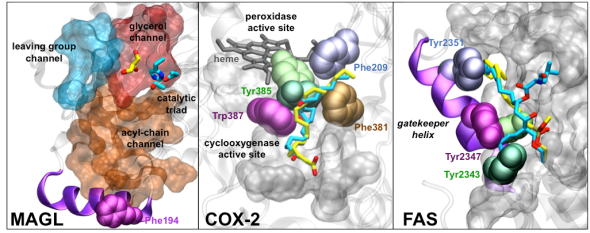
Comparative analysis & experimental validation further reveled an intriguing mechanism of lipid selection (Palermo et al. PLOS Comp. Biol. 2015), in which the interplay between multiple channels, gating residues and substrate flexibility is key for attaining the lipid selection. Based on existing structural data, our results could be extended to other lipid-processing enzymes, which share similar structural features to FAAH. Among them, the endocannabinoid enzyme monoacyl glycerol lipase MAGL,the cyclooxygenases COX-1/2 and the Fatty Acid Synthase FAS enzymes.
We also studied the mechanism of action of covalent inhibitors of FAAH, providing a rational for the activity of novel inhibitors (Palermo et al. J. Med. Chem. 2011; Palermo et al. Eur. J. Med. Chem. 2015). QM/MM calculations revealed that FAAH induces a distortion of the amide bond of cyclic aryl ureas, which definitely favours the formation of a covalent FAAH/inhibitor adduct. These findings could directly help in the rational structure-based design of novel covalent FAAH inhibitors.
Read more: Eur. J. Med. Chem. 2015, 91, pp 15-26
J. Phys. Chem. B. 2015, 119, pp 789–801
J. Chem. Theory. Comput. 2013, 15, pp 1202-1213
J. Med. Chem. 2011, 54, pp 6612–6623